Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
 and Sr
and Sr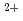 recognition mechanism of halophilic
recognition mechanism of halophilic  -lactamase
-lactamaseArai, Shigeki; Yonezawa, Yasushi; Adachi, Motoyasu; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
Proteins can distinguish various metal ions. Recently, we succeeded in X-ray crystallographic analysis of the halophilic  -Lactamase (HaBLA) derived from
-Lactamase (HaBLA) derived from 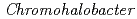 sp.560 under the existence of Cs+ and Sr2+. Diffraction data at 2.0
sp.560 under the existence of Cs+ and Sr2+. Diffraction data at 2.0  resolution (space group P31, Unit cell a = b =115.9
resolution (space group P31, Unit cell a = b =115.9  , c =67.9
, c =67.9  , Rmerge 9.6%) was collected. Three Cs
, Rmerge 9.6%) was collected. Three Cs and six Sr
and six Sr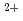 metal ion binding sites of HaBLA molecules in the asymmetric unit were identified. This structural information is very useful to create the artificial Cs
metal ion binding sites of HaBLA molecules in the asymmetric unit were identified. This structural information is very useful to create the artificial Cs or Sr
or Sr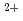 binding site on the protein molecules.
binding site on the protein molecules.
Adachi, Motoyasu; Arai, Shigeki; Matsumoto, Fumiko; Kuroki, Ryota; Hatanaka, Takaaki*; Ito, Yuji*; Hidaka, Koshi*; Tsuda, Yuko*; Kiso, Yoshiaki*
no journal, ,
HIV protease is known as a drug target protein. To compare interactions between HIVPR and inhibitors, single-chained HIVPR linked by a disulfide bond was designed as N98C mutant. Both WT N98C and A17 N98C mutants were expressed as inclusion body and refolded by dilution method. The yield of the purified enzymes was similar to that of WT. We will report the results of interactions between inhibitors and WT N98C and A17 N98C mutants by physicochemical and crystal structure analyses.
Shimizu, Rumi; Adachi, Motoyasu; Kuroki, Ryota; Yamashita, Michi*; Morimoto, Satoshi*
no journal, ,
no abstracts in English
Arimori, Takao; Kawamoto, Noriko*; Okazaki, Nobuo; Nakazawa, Masami*; Miyatake, Kazutaka*; Ueda, Mitsuhiro*; Tamada, Taro
no journal, ,
no abstracts in English
Matsumoto, Fumiko; Hatanaka, Takaaki*; Adachi, Motoyasu; Shimizu, Rumi; Tamada, Taro; Ito, Yuji*; Kuroki, Ryota
no journal, ,
no abstracts in English
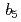 reductase
reductaseYamada, Mitsugu; Tamada, Taro; Matsumoto, Fumiko; Takeda, Kazuki*; Kimura, Shigenobu*; Kuroki, Ryota; Miki, Kunio*
no journal, ,
no abstracts in English
Kurihara, Kazuo; Tamada, Taro; Yamada, Mitsugu; Ohara, Takashi*; Kuroki, Ryota
no journal, ,
no abstracts in English
Higuchi, Mariko; Pinak, M.
no journal, ,
Double strand break (DSB) of DNA is repaired by non-homologous end joining (NHEJ). Repair DSB due to high-LET (linear energy transfer) iron-ion radiation takes longer time than that due to low-LET X rays. DNA damages due to high-LET iron-ion radiation are more condensed than that due to low-LET X rays. Such DSB with other lesions is called complex DSB. The question is why other lesions near DSB cause delay to repair DSB. The first step of NHEJ is considered binding of Ku70/80 to DSB. One reason of delay to repair complex DSB may be inhibition binding of Ku to complex DNA. The aim of this study is clarify if the complex DSB caused inhibition binding of Ku or not. In this study, we present result of molecular dynamics simulation of Ku and DNA including 8oxoG located on interaction region with Ku. The fluctuation of DNA including 8oxoG bound Ku was rather larger than that of native DNA bound Ku. But, binding structural entropy of DNA including 8oxoG was similar as that of native DNA.